The immune system is not only crucial for protection against infectious agents, but plays a key role in responses to tumors and in various chronic diseases. The combination of monoclonal antibodies and flow cytometry has had an enormous impact on immune system studies, but it does not provide information on the spatial distribution of the cells of interest within a tissue. Immunohistochemistry (IHC) provides information about the spatial organization of cells in a tissue, but most IHC methods are of low complexity (2 to 4 colors) insufficient for identifying the many known immune cell types in tissues. The Lymphocyte Biology Section (LBS) of the Laboratory of Immune System Biology (LISB) in NIAID has developed a new IHC approach called Histo-cytometry that involves immunofluorescent staining of tissues in up to 14 colors, imaging at high resolution across large areas, obtaining quantitative information about staining intensity on a per-cell basis, and locating identified cell subsets in a tissue section. Recent improvements include sequential staining to achieve > 30 parameters, addition of in situ mRNA FISH, and development of a new method (Ce3D) that allows applying the Histo-cytometry technique to 3D tissue volumes.
The goal of the Center for Advanced Tissue Imaging (CAT-I), a joint effort of the NIAID Division of Intramural Research and the NCI Center for Cancer Research, is to enable intramural and extramural investigators to collaborate with experts in the Laboratory of Immune System Biology, DIR, NIAID in the application of LBS methods of Histo-cytometry in 2D and 3D, to enable highly multiplex, quantitative image analysis of tissue samples, primarily but not exclusively of human origin, and to use these data to advance our understanding of immune processes in various diseases or to develop better vaccines.
Histo-cytometry
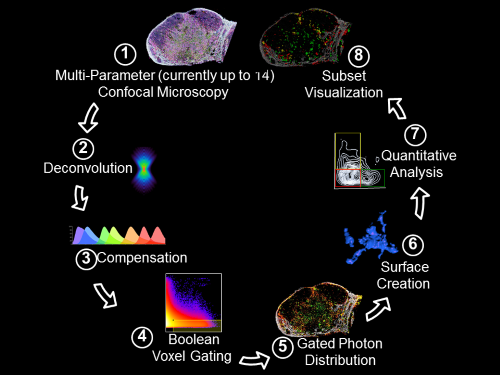
Histo-cytometry workflow pipeline. Gerner et. al., 2012, Immunity 37:364-376
Flow cytometry allows highly quantitative analysis of complex dissociated populations at the cost of neglecting their tissue localization. In contrast, conventional microscopy methods provide spatial information, but visualization and quantification of cellular subsets defined by complex phenotypic marker combinations are challenging. We have developed an analytical microscopy method, Histo-cytometry, for visualizing and quantifying phenotypically complex cell populations directly in tissue sections. This technology is based on multiplexed antibody staining, tiled high-resolution confocal microscopy, voxel gating, volumetric cell rendering, and quantitative analysis. We have tested this technology on various innate and adaptive immune populations in murine, nonhuman primate, and human lymph nodes (LNs) and were able to identify complex cellular subsets and phenotypes, achieving quantitatively similar results to flow cytometry while also gathering cellular positional information.
Ce3D
The spatial organization of cells within tissues is critical for many key biological processes, such as cellular differentiation, intercellular communication, and localized function. Our understanding of cellular positioning has been historically based on tissue section imaging, which discards all three-dimensional (3D) information and is problematic for understanding structure-function relationships of complex organs and relatively rare events. Although recent innovations in tissue clearing methodologies have paved the way for whole organ microscopy, they suffer from various technique-specific artifacts, such as loss of reporter fluorescence, poor epitope preservation, or morphological distortions.
We have developed a new method, Clearing-enhanced 3D (Ce3D), which generates excellent tissue transparency, preserves cellular morphology, and maintains reporter protein fluorescence, while also retaining epitopes for antibody-based labeling. We have used Ce3D imaging to conduct whole-tissue microscopy of various cell types in lymphoid and non-lymphoid organs. When coupled with multiplexed antibody labeling, innovative image segmentation approaches, and Histo-cytometry, Ce3D allows for quantitative analysis of phenotypically complex cells in dense lymphoid organs. Collectively, this methodology provides a no-compromise solution for highly quantitative imaging of intact tissues using conventional confocal microscopes, offering a novel tool for studying cellular organization and position-dependent physiology in complex organs.

3D reconstruction of small intestine.

