The SARS-CoV-2 Interagency Group (SIG), established by the U.S. Department of Health and Human Services (HHS), works to rapidly characterize emerging variants and actively monitors their potential impact on SARS-CoV-2 vaccines, therapeutics, and diagnostics. The SIG is responsible for variant classifications in the United States and meets regularly to evaluate the risk posed by SARS-CoV-2 variants circulating in the United States and globally to make recommendations about the variants. NIAID is a member of the SIG and supports research and surveillance activities to help inform decisions made by the SIG.
To support the generation of data for the SIG, NIAID has created the SARS-CoV-2 Assessment of Viral Evolution (SAVE) risk-assessment pipeline for SARS-CoV-2 variant viruses. SAVE provides a comprehensive real-time risk assessment of emerging mutations in SARS-CoV-2 that could impact transmissibility, virulence, and susceptibility to infection- or vaccine-induced immunity. The goals of the SAVE program are to understand emerging changes in the virus that could impact transmissibility, virulence, susceptibility to convalescent and vaccine-induced immunity and diagnostic testing. SAVE is one of the data streams that provides important information for variant characterization and complements other USG and NIH efforts like the ACTIV Tracking Resistance and Coronavirus Evolution (TRACE) initiative.
The NIAID SAVE program is composed of an international team of scientists with expertise in virology, immunology, vaccinology, structural biology, bioinformatics, viral genetics, and evolution. Researchers from NIAID Intramural, the NIAID Vaccine Research Center, other HHS and Department of Defense laboratories, and the extramural academic community work collaboratively within and across multiple sub-groups to accelerate the pace of variant research and discovery through rapid and open sharing. Researchers are developing high quality data sets that are then vetted by SIG working groups comprised of various experts. These data help inform public health recommendations.
More information on the SAVE program can be found in the publication, "Defining the risk of SARS-CoV-2 variants on immune protection" (Nature, 2022).
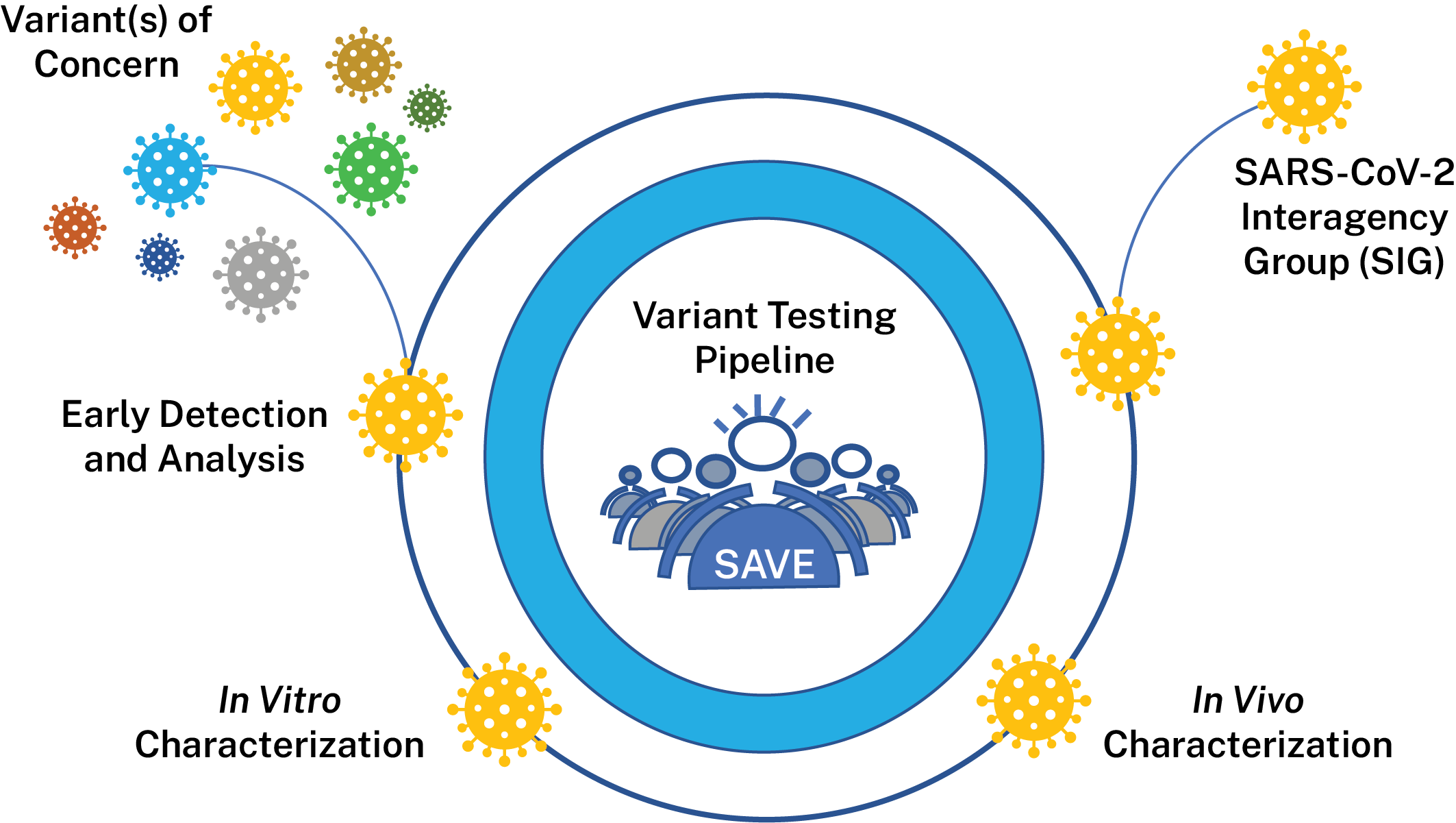
NIAID’s activities are divided into the following data-generating components:
Early Detection and Variant Analysis
Lead: Elodie Ghedin, Ph.D., NIAID and Derek Smith, Ph.D., Cambridge University
This team is evaluating surveillance/epidemiological data, information from observational studies on the efficacy of COVID-19 vaccines, variant information from documented vaccine breakthrough cases and accompanying sequence data from these efforts to identify variants that should be evaluated further in vitro and in vivo. They will also perform antibody escape mapping in the lab using monoclonal antibodies and polyclonal sera in pseudoviral assays to identify proactively substitutions of concern.
In vitro Characterization
Lead: Florian Krammer, Ph.D., Icahn School of Medicine at Mount Sinai and Mehul Suthar, Ph. D., Emory University
This team characterizes variants identified by the early detection group comprehensively in vitro to determine how they may differ from the original SARS-CoV-2 virus. Variant viruses, pseudoviruses and various convalescent and vaccinee sera samples are used to determine if variants are able to evade infection- or vaccine-induced immunity. Groups are also performing antigenic landscape analyses to determine the relatedness of each emerging variant to the original strain to which vaccines were made. Researchers also analyze characteristics such as in vitro replication kinetics to determine how variant viruses may have different characteristics that affect their ability to replicate or transmit between humans. Groups are also exploring if any variants can evade cellular immune responses generated by memory B-cells and T-cells. The data generated by this groups helps determine which variants are of most concern for our existing countermeasures.
In vivo Characterization
Lead: Michael Diamond, M.D., Ph.D., and Jacco Boon, Ph.D., Washington University in St. Louis
This team is using animal models to estimate virulence changes in circulating SARS-CoV-2 viruses and evaluate vaccine efficacy against variant viruses. Researchers use various animal models to examine if any variants cause different levels of disease in models. Researchers also vaccinate or infect animals to determine if animals remain protected from infection with variant viruses over different periods of time. Finally, researchers are working to study ability of countermeasures such as vaccines to prevent transmission in animal models.
Contact Information
Research Resources
SAVE is generating and characterizing reagents to be used across the USG and the global scientific community, including sera panels, viruses and protein. SAVE-generated and other available resources can be found at BEI Resource Repository.

