The Integrated Research Facility at Fort Detrick (IRF-Frederick) team has extensive experience in testing potential medical countermeasures for high-consequence viral pathogens. Leveraging this experience, scientists at the IRF-Frederick have developed and are using cell‑based in vitro assays and animal models to test potential therapeutics for and vaccines against COVID‑19. The unique medical imaging capabilities of the IRF-Frederick are also being used to understand pathological consequences of SARS‑CoV‑2 in relevant animal models of disease. Following are examples of the IRF-Frederick’s contribution to solving the challenge of COVID-19 for the world and for the nation.
High-Throughput Assay Platform
The IRF-Frederick has developed drug-screening and virus-neutralization assays using the PerkinElmer Operetta high-content imaging system. Fluorescence and chemiluminescence cell-based assays provide automated data capture and analysis.
Drug Screening Assay
The IRF-Frederick is currently working with both government and industry partners to screen individual compounds and combinations of compounds in cell-based high-content imaging assays.
The IRF-Frederick has developed cell-culture screening assays to identify potentially effective therapeutics for Ebola, Marburg, Lassa, and Nipah viruses, MERS-CoV and SARS-CoV-2.
Microneutralization Assay
Antibody-based therapies demonstrate potential as treatment for SARS-CoV-2 infection. NIAID’s Division of Clinical Research (DCR) worked with commercial manufacturers of intravenous immunoglobulin (IVIG) for use in in-patient and out-patient clinical trials (e.g., NCT04546581). IRF Frederick scientists used a microneutralization assay to screen donor plasma and therapeutic antibody products. This assay is also being used to test a range of monoclonal antibodies, plasma, nanobodies, and purified IgG as potential therapeutics in support of multiple collaborators, including intramural and extramural researchers.
Immunoglobulin M (IgM) and Immunoglobulin G (IgG) Enzyme-Linked Immunosorbent Assays (ELISAs)
As part of in-house experimental animal-model work, the IRF-Frederick scientists developed IgM and IgG ELISAs for SARS-CoV-2, targeting both the viral spike protein and the receptor-binding domain (RBD) within the spike protein. The SARS-CoV-2 assays are being developed for both animal and human studies to evaluate the efficacy of medical countermeasures, particularly vaccines.
Small Animal Model Development
IRF-Frederick scientists are using golden hamsters as the disease model of choice for COVID-19 studies. Exposure by intranasal inoculation of SARS-CoV-2 results in mild to moderate disease in hamsters, including weight loss, measurable viral loads in the lungs and respiratory tract, and lung pathology similar to that observed in COVID-19 patients. Other disease parameters measured in studies are the presence of viral RNA in the upper and lower respiratory tracts, extrapulmonary organs, and blood harvested from virus-infected hamsters. Histopathological analysis of lung samples identified interstitial pneumonia on Day 2 and Day 5, with resolution at Day 8 post-exposure. While the hamster has proven a very effective model for most SARS-CoV-2 variants identified to date, the omicron variant (B.1.1.529) causes a very mild disease that is not clinically apparent in this model and requires determination of viral loads in the upper respiratory tract to measure disease. The hamster model has been used to evaluate multiple potential therapeutic products for efficacy against several SARS-CoV-2 variants, including omicron.
Capabilities
- Aerosol exposure of small animal models
- Use of positron emission tomography (PET), computed tomography (CT) imaging, and fluorodeoxyglucose (FDG) PET/CT to monitor disease progression in vivo and longitudinally, with specific focus on lung consolidation and inflammation
- Testing of medical countermeasures, including vaccine candidates, antiviral drugs, and antibody therapeutics against SARS-CoV-2
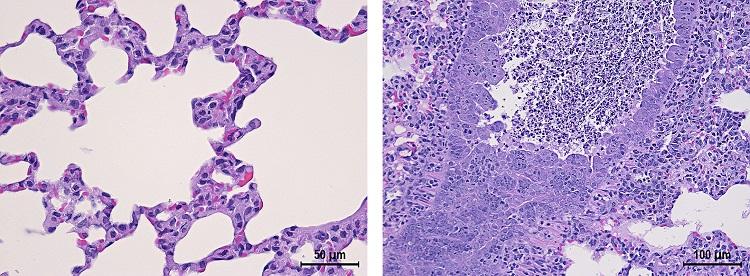
Representative histopathology from mock-infected control or infected hamsters indicating interstitial pneumonia similar to disease observed in humans: control (left) and 106 PFU (right).
The IRF-Frederick has developed imaging to assess [fluorine-18]fluorodeoxyglucose (18F-FDG) kinetic changes in the lungs of aged golden hamsters versus young in the context of evaluating disease progression. Aged hamsters develop more severe lung disease than young hamsters. Interstitial pneumonia was demonstrated and quantified through imaging, which helps overcome limitations of these models by providing the following results:
- Qualitative CT assessments that mimic human disease
- Quantitative CT assessments that correspond to histopathology
- PET signal corresponds to CT lung lesions
- Gross and histopathology support radiological findings
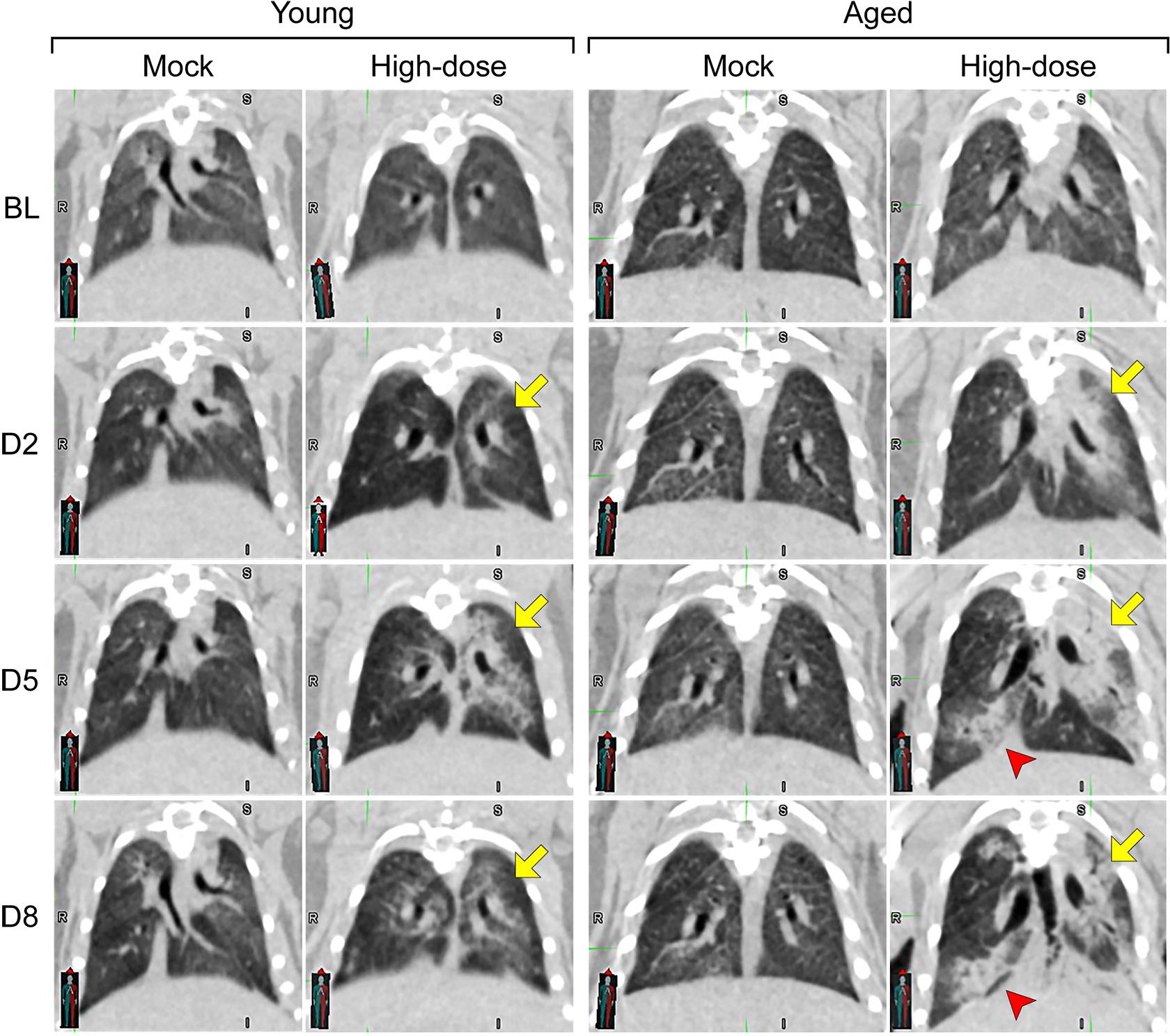
Representative computed tomography (CT) images from mock and high-dose groups of young (4 to 5 weeks) and aged (1 year) golden hamsters: From top to bottom, images show disease progress (baseline [BL] to Day 8). The two columns on the left are lung images from young hamsters, and the two on the right are from aged hamsters. In the high-dose group, disease peaked in young hamsters on Day 5 and resolved on Day 8, while aged hamsters showed mixed responses on Day 8 (yellow and red arrows). Both young and aged hamsters that received a mock dose had normal lungs, while hamsters infected with the high dose showed ground-glass opacities and consolidation, which mimic human disease.
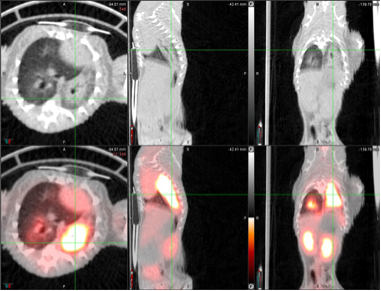
Uptake of [fluorine-18]fluorodeoxyglucose (18F-FDG), quantified by positron emission tomography (PET), (generally) correlates with lung consolidation detected by computed tomography (CT).
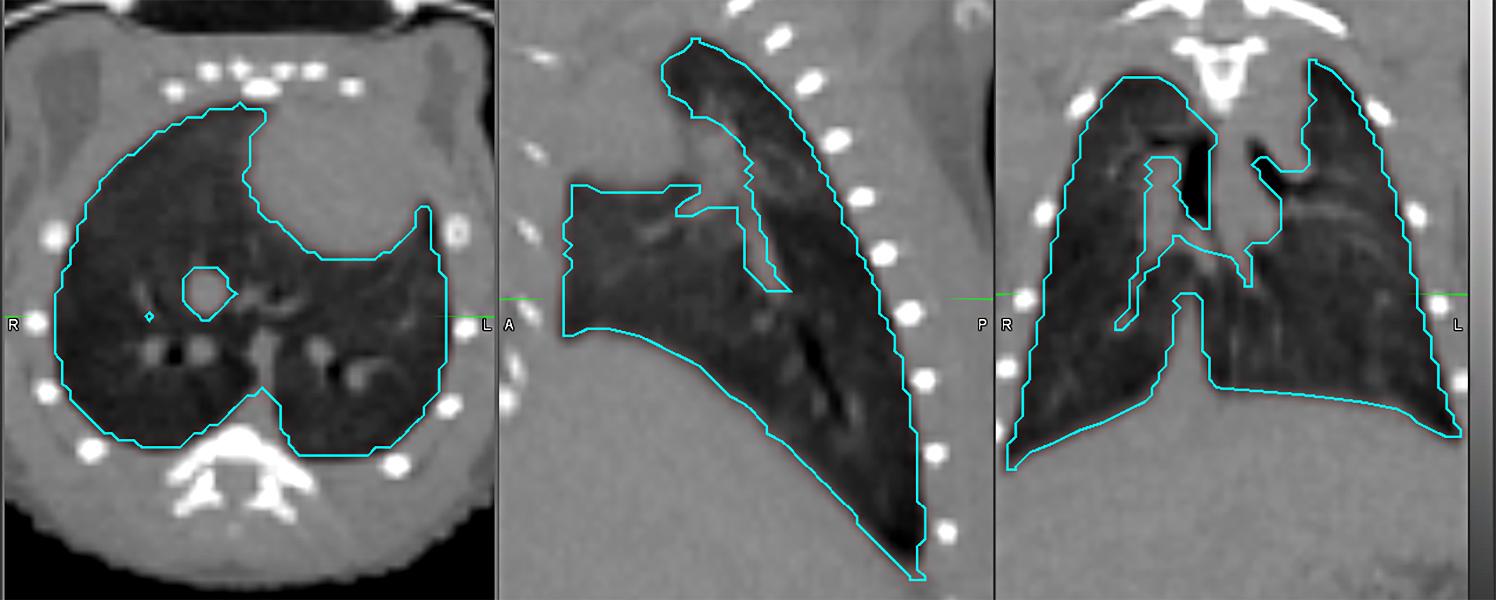
Semi-automatic lung segmentation for computed tomography (CT) scoring: axial (left), sagittal (center), and coronal (right) views.
Nonhuman Primate Model Development
IRF-Frederick scientists have developed a nonhuman primate model through a natural history study of SARS-CoV-2 in crab-eating (cynomolgus) macaques. Nonhuman primate studies at the IRF‑Frederick include a variety of SARS-CoV-2-specific assays, as well as CT imaging and percent change in lung hyperdensity (PCLH), to quantifiably and objectively evaluate disease progression and the efficacy of medical countermeasures. The unique medical imaging capabilities of the IRF‑Frederick enable the exploration of correspondence between organ structure (CT scan) and physiology (fluorodeoxyglucose [FDG] PET).
Three-dimensional rendering of crab-eating macaque (Macaca fascicularis) lungs from qualitative computed tomography (CT) before and after intrabronchial instillation of SARS-CoV-2 (from baseline [BL] to Day 6 [D6]); airways (blue); normal lung (gray); vessels (red); imaging abnormalities (yellow). See more SARS-CoV-2 imaging examples.
Credit: NIAID
Capabilities
- Assays
- Real-time reverse transcription polymerase chain reaction (RT-qPCR)
- Fluorescence reduction neutralization assay (FRNA)
- Plaque assay
- In situ hybridization (ISH), immunohistochemistry (IHC), and IgG ELISAs for total spike protein (total S), N‑terminal globular subunit 1 (S1), and subunit 2 (S2)
- Imaging
- CT
- Percent change in lung hyperdensity (PCLH)
- FDG PET
Human Clinical Trials
The Clinical Studies Support Team (CSST) at the IRF-Frederick is actively engaged in the design of and support for multiple clinical studies. CSST members have assisted the Partnership for Research on Emerging Viruses in Liberia (PREVAIL) with the development and implementation of a research study to characterize the clinical course, rate of disease progression, risk factors for progression, and death from SARS-CoV-2 infections. The study includes long-term follow-up and exploration of the risks of re-infection and immunologic responses to SARS-CoV-2. In addition, CSST is supporting International Study on COVID-19 Vaccine to Assess Immunogenicity, Reactogenicity and Efficacy (InVITE) in Liberia, Guinea, and the Democratic Republic of the Congo to study COVID-19 vaccine immunogenicity and durability, as well as SARS-CoV-2 infections in people who received an initial vaccine or booster vaccine regimen. The team is also supporting the international Adaptive COVID-19 Treatment Trial (ACTT) by preparing critical laboratory reagents and aiding in the establishment of sequencing capabilities to monitor emergence of variants.

