Structural Bioinformatics Core
Tongqing Zhou, Ph.D.
Contact: tzhou@mail.nih.gov

Major Areas of Research
- Serum Analysis/Antibodyomics
- Immunogen Design
- Structural Bioinformatics
Program Description
The Structural Bioinformatics Core (SBIS) seeks to apply the tools of computational biology and structural bioinformatics to the design of an effective HIV-1 vaccine. Over the last few years, these tools have met with growing success when applied to a wide range of problems, including protein design, protein-structure prediction, enzyme design, and drug design. Our goal is to utilize available state-of-the-art structural bioinformatics tools, as well as to develop novel methodologies, as part of a collaborative effort—within the Vaccine Research Center, with other intramural portions of the National Institutes of Health, and extramurally—to assist in development of effective vaccines against HIV-1 and other viruses.
The efforts of the SBIS can be divided into three areas, graphically shown here and explained below.
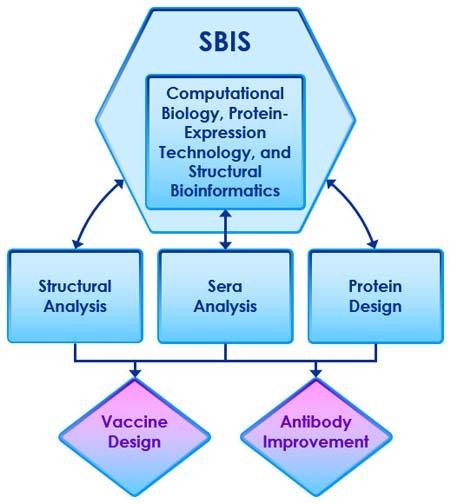
The three areas of Structural Bioinformatics Core (SBIS): Computational Biology, Protein-Expression Technology, and Structural Bioinformatices
Bioinformatics and Vaccine Design
1. Using computational tools to analyze structure and function. The solution of crystal structures and in-depth structural analysis play a pivotal role in current efforts for rational immunogen design. Often, a significant amount of structural information exists about an important biological system (e.g., for protein subunits or from cryo-electron tomography), though the central biological target resists atomic-level structural analysis (e.g., the functional viral spike of HIV-1). Computational biology can serve as a bridge between these other sources of information and the design of appropriate crystallization constructs, to enable atomic-level analysis of a particular target. Moreover, once the structure of a particular target is determined, computational biology can assist in the analysis of the structure, to extract biological meaning.
2a. Computationally assisted analysis of sera and isolation of monoclonal antibodies. An understanding of the serum responses of both HIV-1-infected individuals and vaccines should assist in the development of an effective HIV-1 vaccine. Computational design can assist in the development of antigenically specific probes useful in analyzing the neutralizing activity of sera; computation analysis can also assist in deciphering the HIV-1 elements recognized by both binding and neutralizing antibodies in sera. Such an understanding provides critical in vivo feedback for the iterative structure-based improvement of immunogens.
2b. Antibodyomics: bioinformatics of next-generation sequencing of B cell transcripts. A critical aspect of vaccine design is the development of B cells that produce antibodies capable of neutralizing virus. Currently, however, little is understood about how antibodies develop in response to viruses such as HIV-1. To that end, next-generation sequencing of B cell transcripts provides a wealth of information on the sequences of antibody heavy and light chains—a single dataset may contain up to hundreds of thousands of antibody sequences. Mining such large datasets for relevant information is non-trivial, and the development and application of bioinformatics tools for the analysis of next-generation antibody sequencing data is an essential step in the quest for understanding the development of B cell lineages, which could provide clues about pathways to the design of successful vaccines.
Recent progress in the field has provided encouragement that development of a vaccine for HIV-1 is possible. By combining the power and efficiency of computation with the wealth of information encoded in protein structures, SBIS can play a central role in the vaccine design process.
3. Application of computational techniques to structure-based immunogen design. Specifically, a variety of techniques can be used to focus the immune response toward target epitopes and away from undesirable, often immunoprominent regions, through an iterative process of structure-based design, immunogenic evaluation, computational manipulation, and immunogen redesign. This process takes advantage of other skill sets resident within the Virology Laboratory, specifically of the ability of the Structural Biology Core to provide atomic-level details on the target epitope and of the Vector core to evaluate immunogens. We expect this strategy of rational immunogen design to lead to the elicitation of antibodies that broadly neutralize a diverse range of HIV-1 isolates. The direct rational structure-based design of antibodies is also of interest to the SBIS.
Biography
Education
Ph.D., Chinese Academy of Sciences, Beijing, China
M.Sc., Wayne State University, Detroit, MI
Dr. Tongqing Zhou joined the Dale and Betty Bumpers Vaccine Research Center in 2001, became a staff scientist in 2005, and was promoted to chief of the NIAID Structural Bioinformatics Core in 2018. Dr. Zhou received his Ph.D. in cell biology from the Chinese Academy of Sciences and his M.Sc. in electronic and computer controlled systems from Wayne State University. He is joined by the two co-heads of the core, Drs. Gwo-Yu Chuang and Chen-Hsiang Shen. Dr. Gwo-Yu Chuang received his Ph.D. training under the mentorship of Dr. Sandor Vajda, Boston University. Dr. Shen received his Ph.D. training under the mentorship of Dr. Irene Weber, Georgia State University. Drs. Peter Kwong and Lawrence Shapiro also serve as advisory mentors to the Structural Virology and Vaccinology Section.
Selected Publications
Mukhamedova M, Wrapp D, Shen CH, Gilman MSA, Ruckwardt TJ, Schramm CA, Ault L, Chang L, Derrien-Colemyn A, Lucas SAM, Ransier A, Darko S, Phung E, Wang L, Zhang Y, Rush SA, Madan B, Stewart-Jones GBE, Costner PJ, Holman LA, Hickman SP, Berkowitz NM, Doria-Rose NA, Morabito KM, DeKosky BJ, Gaudinski MR, Chen GL, Crank MC, Misasi J, Sullivan NJ, Douek DC, Kwong PD, Graham BS, McLellan JS, Mascola JR. Vaccination with prefusion-stabilized respiratory syncytial virus fusion protein induces genetically and antigenically diverse antibody responses. Immunity. 2021 S1074-7613(21)00116-3.
Rawi R, Rutten L, Lai YT, Olia AS, Blokland S, Juraszek J, Shen CH, Tsybovsky Y, Verardi R, Yang Y, Zhang B, Zhou T, Chuang GY, Kwong PD, Langedijk JPM. Automated Design by Structure-Based Stabilization and Consensus Repair to Achieve Prefusion-Closed Envelope Trimers in a Wide Variety of HIV Strains. Cell Report. 2020 33(8):108432.
Shen CH, DeKosky BJ, Guo Y, Xu K, Gu Y, Kilam D, Ko SH, Kong R, Liu K, Louder MK, Ou L, Zhang B, Chao CW, Corcoran MM, Feng E, Huang J, Normandin E, O'Dell S, Ransier A, Rawi R, Sastry M, Schmidt SD, Wang S, Wang Y, Chuang GY, Doria-Rose NA, Lin B, Zhou T, Boritz EA, Connors M, Douek DC, Karlsson Hedestam GB, Sheng Z, Shapiro L, Mascola JR, Kwong PD. VRC34-Antibody Lineage Development Reveals How a Required Rare Mutation Shapes the Maturation of a Broad HIV-Neutralizing Lineage. Cell Host Microbe. 2020 27(4):531-543.e6.
Joyce MG, Wheatley AK, Thomas PV, Chuang GY, Soto C, Bailer RT, Druz A, Georgiev IS, Gillespie RA, Kanekiyo M, Kong WP, Leung K, Narpala SN, Prabhakaran MS, Yang ES, Zhang B, Zhang Y, Asokan M, Boyington JC, Bylund T, Darko S, Lees CR, Ransier A, Shen CH, Wang L, Whittle JR, Wu X, Yassine HM, Santos C, Matsuoka Y, Tsybovsky Y, Baxa U; NISC Comparative Sequencing Program, Mullikin JC, Subbarao K, Douek DC, Graham BS, Koup RA, Ledgerwood JE, Roederer M, Shapiro L, Kwong PD, Mascola JR, McDermott AB.Vaccine-Induced Antibodies that Neutralize Group 1 and 2 Influenza A Viruses.Cell. 2016 166(3):609-23.
Zhou T, Zhu J, Wu X, Moquin S, Zhang B, Acharya P, Georgiev IS, Altae-Tran HR, Chuang GY, Joyce MG, Do Kwon Y, Longo NS, Louder MK, Luongo T, McKee K, Schramm CA, Skinner J, Yang Y, Yang Z, Zhang Z, Zheng A, Bonsignori M, Haynes BF, Scheid JF, Nussenzweig MC, Simek M, Burton DR, Koff WC; NISC Comparative Sequencing Program, Mullikin JC, Connors M, Shapiro L, Nabel GJ, Mascola JR, Kwong PD.Multidonor analysis reveals structural elements, genetic determinants, and maturation pathway for HIV-1 neutralization by VRC01-class antibodies.Immunity. 2013 39(2):245-58.
Wu X, Zhou T, Zhu J, Zhang B, Georgiev I, Wang C, Chen X, Longo NS, Louder M, McKee K, O'Dell S, Perfetto S, Schmidt SD, Shi W, Wu L, Yang Y, Yang ZY, Yang Z, Zhang Z, Bonsignori M, Crump JA, Kapiga SH, Sam NE, Haynes BF, Simek M, Burton DR, Koff WC, Doria¬Rose N, Connors M; NISC Comparative Sequencing Program, Mullikin JC, Nabel GJ, Roederer M, Shapiro L, Kwong PD, Mascola JR.Focused Evolution of HIV¬1 Neutralizing Antibodies Revealed by Structures and Deep Sequencing.Science. 2011 333(6049):1593¬602.
Research Group

Top row from left to right: Lawrence Shapiro, Chen-Hsiang Shen
Second row: Tatsiana Bylund, Reda Rawi
Third row: Margarita Mejia, Katie Farney, Baoshan Zhang, Gwo-Yu Chuang
Bottom row: Peter Kwong, Yongping Yang

