Molecular Pathogenesis Section
Established in 2019
Emmie de Wit, Ph.D.
Senior Investigator, Molecular Pathogenesis Section

Major Areas of Research
- Pathogenesis of emerging viruses that cause severe respiratory disease
- Develop human organoid models and animal models to integrate analyses of pathogen, single cell, and host to identify common pathways involved in disease progression
- Study emerging respiratory virus infections in the upper and lower respiratory tract and in the central nervous system
- Use our knowledge of the pathogenesis of respiratory viruses to aid the development of effective, broad-acting therapeutics
Program Description
The goal of the Molecular Pathogenesis Section (MPS) is to increase our understanding of the pathogenesis of emerging respiratory viruses on every level, from host to molecule. The emergence of SARS-CoV-2 in late 2019 and the resulting COVID-19 pandemic have made clear how little we understand about the pathogenesis of viral infections of the respiratory tract and how inadequate existing therapeutics are.
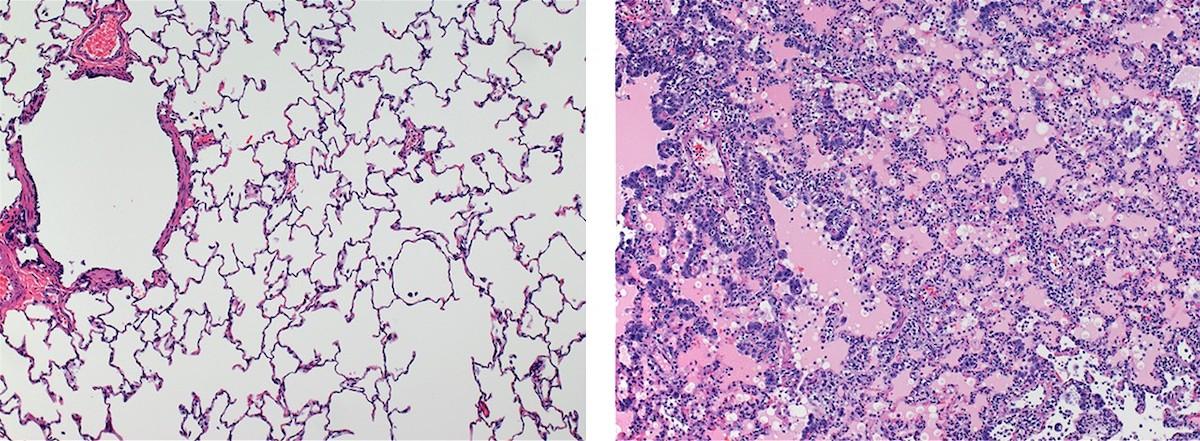
Comparison of healthy lungs (left) to lungs affected by interstitial pneumonia during acute virus infection (right).
It is important to note that even before the COVID-19 pandemic, lower respiratory tract infections were the leading cause of infectious disease deaths worldwide and the eighth most important cause of death in the US. Besides this continuous burden of endemic respiratory viruses, emerging respiratory viruses remain an important threat to global health. Novel respiratory viruses keep emerging from animal reservoirs: in the last decade alone, Middle East Respiratory Syndrome-coronavirus (MERS-CoV), enterovirus D68, Nipah virus, several avian influenza viruses, and SARS-CoV-2 emerged or re-emerged.
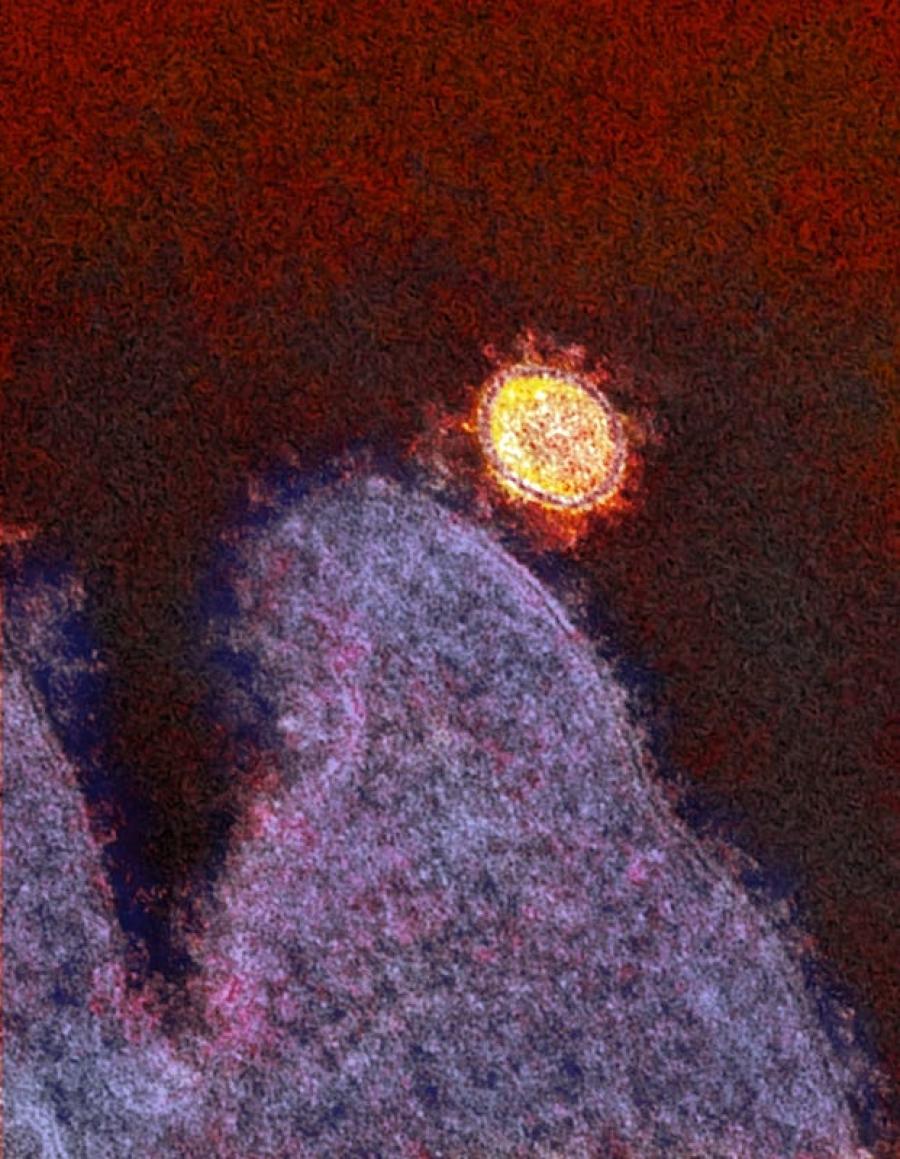
Colorized electron micrograph of a SARS-CoV-2 virus particle in the lungs of an infected rhesus macaque.
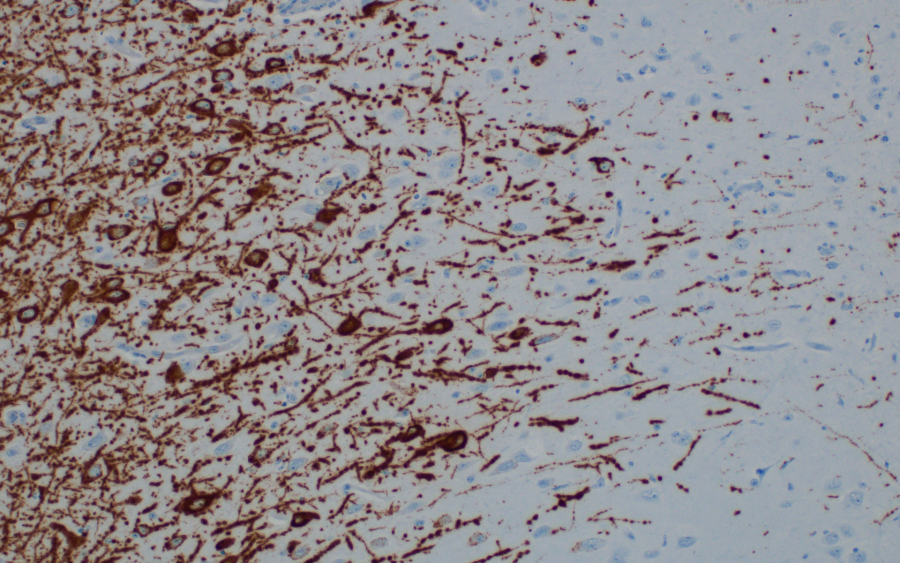
Nipah virus RNA detected by in situ hybridization in neurons in the brain of an African green monkey infected with Nipah virus.
Although great progress has been made in the field of direct-acting antivirals, especially for the treatment of COVID-19, therapeutics to treat severe lower respiratory tract disease have lagged behind. A key problem in the development of treatments for severe respiratory virus disease is that virus replication often peaks ahead of disease severity. Rather than virus-induced lung damage alone, the proinflammatory immune response also contributes to severe respiratory disease. Although increasing attention is given to the host processes involved in severe respiratory infections, these studies are often hampered by a focus on in vivo pathogenesis in lethal disease models, or on mechanistic studies of single molecules or signaling pathways in vitro. In MPS, we aim to combine pathogenesis studies with detailed molecular analyses to identify molecular determinants of severe respiratory tract disease within the virus and the host. Ultimately, the identification of common pathways involved in lower respiratory tract disease progression and druggable targets within those pathways will be used to develop broad-acting, syndrome-based therapeutics.
Many emerging viruses that cause respiratory disease also cause neurological symptoms, e.g., COVID-19, influenza A virus, and Nipah virus. MPS is studying the neuropathogenesis of emerging respiratory viruses in human cerebral organoids and animal models. Currently, our focus is mainly on the neuropathogenesis of Nipah virus.
Biography
Education
Ph.D., Erasmus University Rotterdam, Netherlands
Dr. de Wit received her Ph.D. in virology from Erasmus University Rotterdam, the Netherlands. Her research there focused on the replication, pathogenesis, and transmission of influenza A virus. Dr. de Wit then moved to the Laboratory of Virology of NIAID in Hamilton, Montana, to work in the biosafety level 4 laboratory there. Here, she focused on the pathogenesis of and countermeasures against Nipah virus, the Middle East Respiratory Syndrome Coronavirus and the 1918 H1N1 influenza A virus (Spanish flu). In 2014-2015, Dr. de Wit spent 4 months in a field lab in Monrovia, Liberia, in charge of patient diagnostics for several Ebola Treatment Units in the area to help contain the devastating Ebola epidemic in Liberia. In 2019, Dr. de Wit became the Chief of the Molecular Pathogenesis Section. When SARS-CoV-2 emerged in late 2019, Dr. de Wit focused her research on SARS-CoV-2, developing animal models and using those for testing of medical countermeasures and gaining a better understanding of SARS-CoV-2 pathogenesis. Amongst other accomplishments, the data generated in Dr. de Wit’s lab contributed to the licensing of remdesivir as an antiviral treatment for COVID-19 patients.
Clinical Studies
Nasal Epithelium Samples From Healthy Volunteers
Establishment of Biobank of Nasal Epithelium Samples From Healthy Volunteers
- ClinicalTrials.gov ID NCT06120244
- Keywords: Nasal Epithelium
Selected Publications
Flagg M, Williamson BN, Ortiz-Morales JA, Lutterman TR, de Wit E. Comparison of Contemporary and Historic Highly Pathogenic Avian Influenza A(H5N1) Virus Replication in Human Lung Organoids. Emerg Infect Dis. 2024 Jan 8;31(2).
Singh M, Goldin K, Flagg M, Williamson BN, Lutterman T, Smith B, de Wit E. Intracranial inoculation rapidly induces Nipah virus encephalitis in Syrian hamsters. PLoS Negl Trop Dis. 2024 Oct 28;18(10):e0012635.
Flagg M, Goldin K, Pérez-Pérez L, Singh M, Williamson BN, Pruett N, Hoang CD, de Wit E. Low level of tonic interferon signalling is associated with enhanced susceptibility to SARS-CoV-2 variants of concern in human lung organoids. Emerg Microbes Infect. 2023 Dec;12(2):2276338.
Speranza E, Purushotham JN, Port JR, Schwarz B, Flagg M, Williamson BN, Feldmann F, Singh M, Pérez-Pérez L, Sturdevant GL, Roberts LM, Carmody A, Schulz JE, van Doremalen N, Okumura A, Lovaglio J, Hanley PW, Shaia C, Germain RN, Best SM, Munster VJ, Bosio CM, de Wit E. Age-related differences in immune dynamics during SARS-CoV-2 infection in rhesus macaques. Life Sci Alliance. 2022 Jan 17;5(4):e202101314.
Williamson BN, Feldmann F, Schwarz B, Meade-White K, Porter DP, Schulz J, van Doremalen N, Leighton I, Yinda CK, Pérez-Pérez L, Okumura A, Lovaglio J, Hanley PW, Saturday G, Bosio CM, Anzick S, Barbian K, Cihlar T, Martens C, Scott DP, Munster VJ, de Wit E. Clinical benefit of remdesivir in rhesus macaques infected with SARS-CoV-2. Nature. 2020 Sep;585(7824):273-276.
Munster VJ, Koopmans M, van Doremalen N, van Riel D, de Wit E. A Novel Coronavirus Emerging in China - Key Questions for Impact Assessment. N Engl J Med. 2020 Feb 20;382(8):692-694.
Research Group
The goal of the Molecular Pathogenesis Section (MPS) is to increase our understanding of the pathogenesis of emerging respiratory viruses on every level, from host to molecule. We study the pathogenesis of SARS-CoV-2, Nipah virus, and the highly pathogenic avian influenza H5N1 virus in animal models and human lung and brain organoids.


