Molecular Biology Section
David H. Margulies, M.D., Ph.D.
Chief, Molecular Biology Section

Major Areas of Research
- MHC class I and class II molecules, whose function is to present antigens to T lymphocytes
- Viral immunoevasins and related molecules, in particular those encoded by cytomegaloviruses, that mimic MHC-I molecules in structure and function to modulate the immune response as decoy receptors or by other mechanisms
- Immune hypersensitivity reactions related to MHC-I molecules
- Natural killer (NK)-cell receptors, cell surface molecules of effector cells of the innate immune system that mediate recognition of tumor- and virus-infected cells via the level and composition of MHC-I molecules on the NK-cell target
- T-cell receptors, which by clonal expression confer antigen and MHC specificity for the activation of T cells
Program Description
The Molecular Biology Section (MBS) is directed toward a precise molecular understanding of critical interactions that initiate, control, and perpetuate the immune response. To this aim, the laboratory focuses on:
- The structure and function of cell surface molecules encoded by the major histocompatibility complex (MHC), both the MHC class I and class II molecules
- Function and structure of chaperones involved in peptide loading of MHC I molecules, specifically tapasin and TAPBPR
- Viral immunoevasins related to the MHC-I molecules, particularly those encoded by the cytomegaloviruses
- Immune hypersensitivity reactions related to the expression of specific MHC-I molecules, in particular the role of the human MHC-I molecule HLA-B*57:01 in hypersensitivity to the antiretroviral drug, abacavir
- T-lymphocyte receptors (TCR), which by clonal expression confer antigen and MHC specificity for the activation of T lymphocytes
- Natural-killer (NK)-cell receptors, cell surface molecules of effector cells of the innate immune system, that mediate recognition of tumor and virally infected cells via the level and composition of MHC-I molecules on the NK cell target
- Structure, function, and recognition properties of antibodies directed against SARS-CoV-2, with a focus on X-ray studies of single chain antibodies
- Engineering of high avidity and biparatopic variants of anti-SARS antibodies
In recent years, the major accomplishments of MBS have included determination of the structure and function of catalytic chaperones that contribute to the process of antigen presentation, with particular emphasis on the TAPBPR (TAP binding protein, related), and tapasin, a crucial component of the peptide loading complex (PLC). Using purified MHC molecules, chaperones, viral immunoevasins, TCR, and NK receptors, we have learned the thresholds of affinity that allow such cell surface receptors to activate or inhibit effector-cell function. Based on X-ray crystallographic and nuclear magnetic resonance (NMR) studies of these components and of their molecular complexes, we now understand in molecular detail the mechanism of peptide loading of MHC-I molecules, the basis of molecular recognition of TCR for MHC-I and MHC-II molecules, and the structural basis for interaction of T-cell coreceptors with MHC molecules. X-ray structures of MHC-II/peptide complexes that are crucial to the development of autoimmunity provide insight into the molecular basis of autoimmunity.
Our studies of viral immunoevasins reveal the structural evolution of viral resistance to recognition by the immune system and provide opportunities for developing strategies to counter such viral evasion. In addition, our broad understanding of the structure-function relationships of MHC molecules has permitted us to explore the relationship of MHC-I structure to HLA-linked drug-induced hypersensitivity syndromes. We have developed mouse model systems to allow further exploration of the molecular and cellular basis of such HLA-linked syndromes. In addition, collaborative studies demonstrate the potential use of pan anti-MHC-I antibodies in activating the immune response as a therapy for viral infection or tumors. Finally, structural studies of several synthetic nanobodies (sybodies) directed to SARS-CoV-2 form the basis for ongoing studies to design high-affinity anti-SARS reagents.
The Figures below illustrate several structures determined in the laboratory, that of the tapasin homolog, TAPBPR, bound to its MHC-I substrate, H2-Dd (PDB ID: 5WER, Figure 1); that of sybodies Sb68 and Sb45 bound to the SARS-CoV-2 receptor binding domain (RBD) (PDB ID: 7KLW, Figure 2). Figure 3 illustrates the binding affinity determination of these two sybodies to the wild type (WT) and several mutant RBDs.
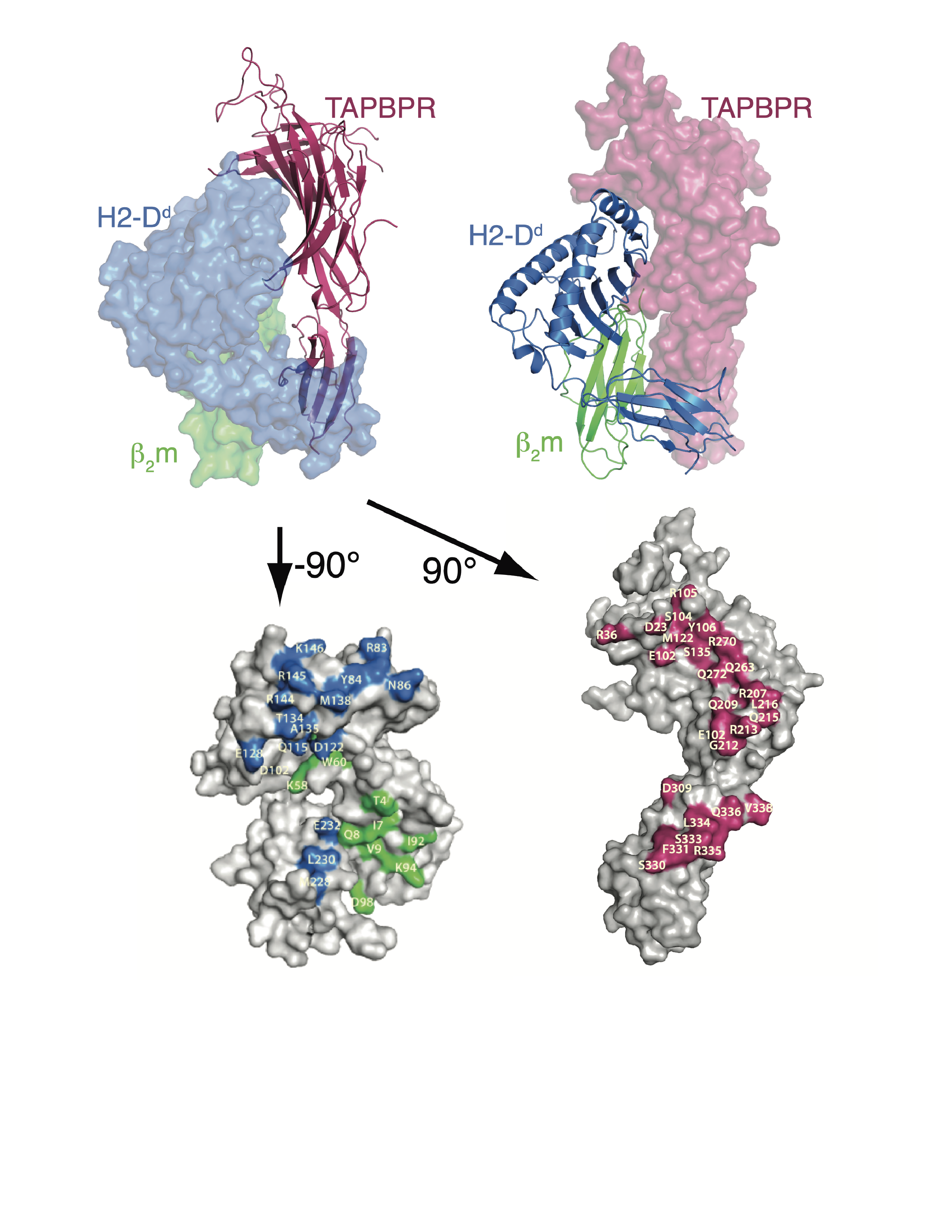
X-ray structure of the MHC-I chaperone, tapasin homolog, TAPBPR (magenta), complexed with the MHC-I molecule, H2-Dd (blue), and its light chain, 2-microglobulin (blue). (Jiang et al, 2017)
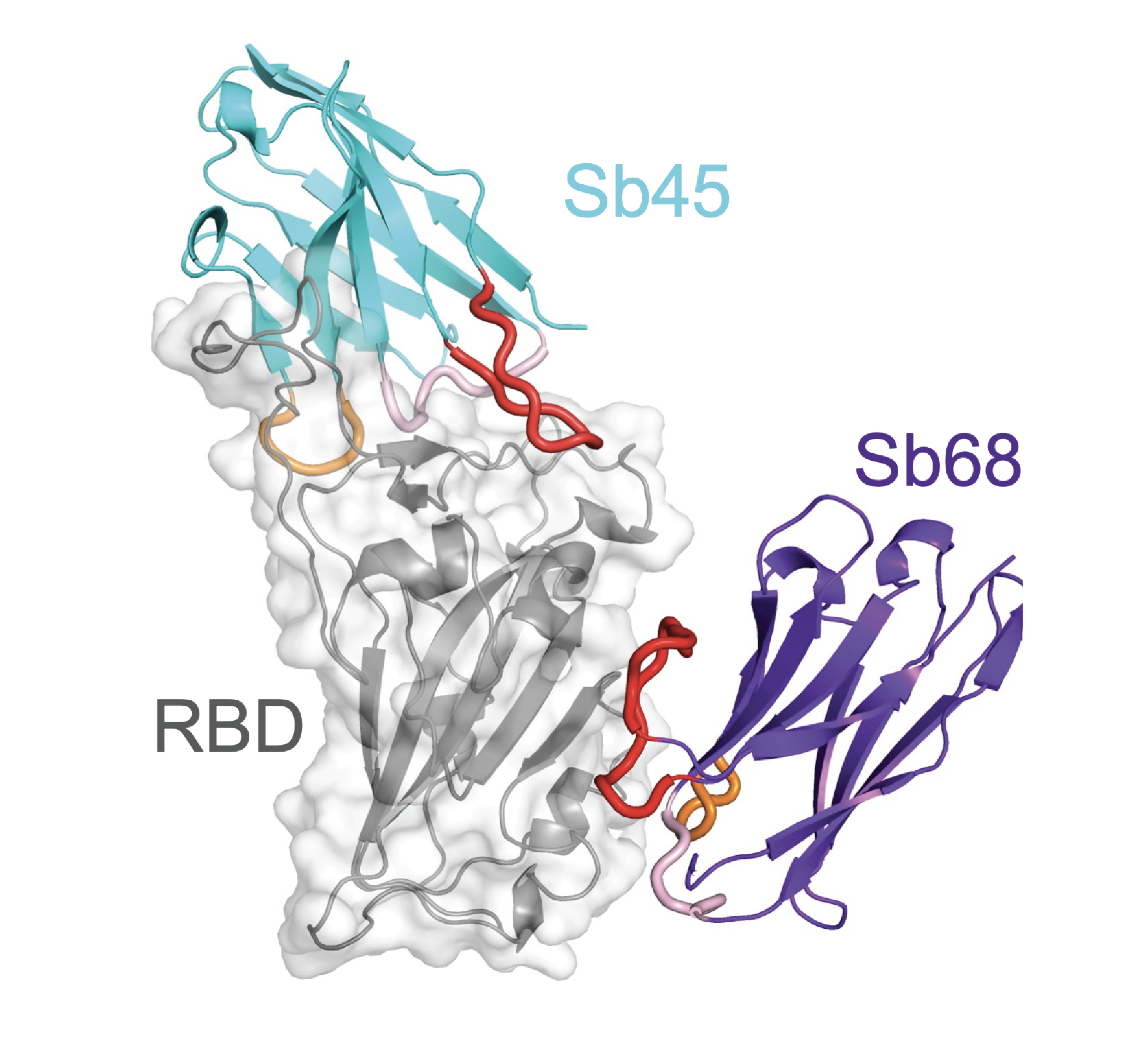
Figure 2. X-ray structure of two single-chain synthetic antibodies (sybodies) bound to the receptor binding domain (RBD) of SARS-CoV-2. (WT, inset) (Ahmad et al. 2021)
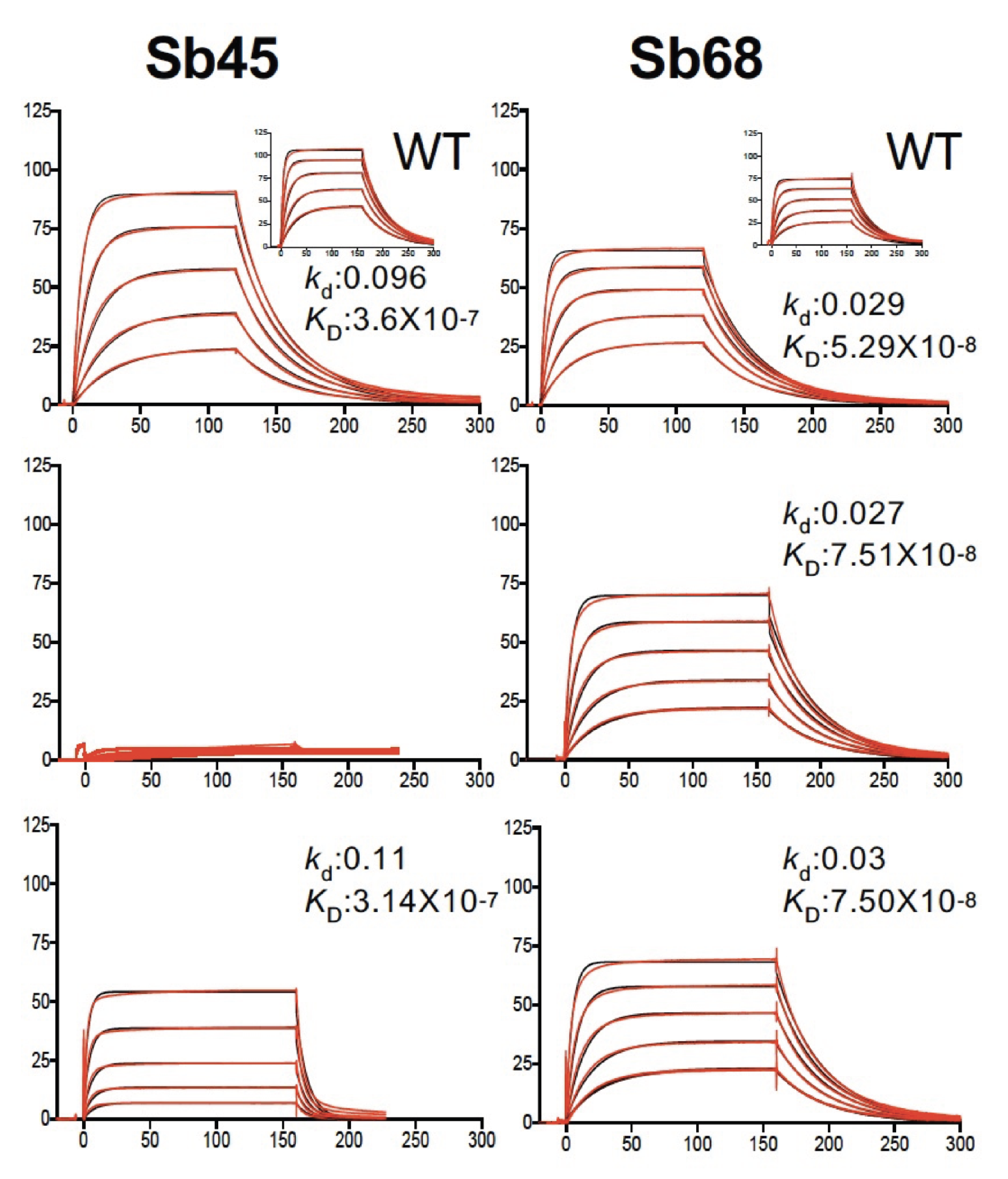
Figure 3. Binding studies of two sybodies to the wild type (WT, inset) and three point mutants of SAS-CoV-2 show sensitivity of Sb45 to the E484K mutation (Ahmad et al 2021)
Biography
Education
M.D., Ph.D., 1978, Albert Einstein College of Medicine
Dr. Margulies received an A.B. from Columbia University in 1971. In 1978, he earned his M.D. and Ph.D. from the Albert Einstein College of Medicine. From 1978 to 1980, he served as a resident in medicine at Columbia/Presbyterian Medical Center. From 1980 to 1983, he worked as a research associate in the Laboratory of Molecular Genetics at the National Institute of Child Health and Human Development. From 1983 to 1987, he was an investigator in the Laboratory of Immunology. In 1987, he became a senior investigator and, since 1989, has been chief of the Molecular Biology Section. Since 2008, he has been a member of the Senior Biomedical Research Service.
Awards
Phi Beta Kappa; U.S. Public Health Service Commendation Medal, 1987; Outstanding Service Medal, 1991; Meritorious Service Medal, 1997; Distinguished Service Medal, 2001; NIH Merit Award, 2009 (Tetramer Facility Team)
Memberships
- The American Association of Immunologists
- American Society for Clinical Investigation
- American Association for the Advancement of Science
- Federation of American Scientists
- Protein Society
Advisory/Review Committees
- Immunology and Immunotherapy Advisory Committee (American Cancer Society, 1988-1991)
- Fellowship Subcommittee (Arthritis Foundation, 1989-1991)
- NIH Research Scholars Program Committee (Howard Hughes Medical Institute, 1991-1993)
- National Multiple Sclerosis Society Advisory Committee, 1993-1999
- The American Association of Immunologists, Publications Committee, 1995-1999
- Department of Veterans’ Affairs Joint Biomedical Development Scientific Merit Review Board, Subcommittee for Immunology-B, 2005-2009
- NIH Immunology Interest Group Steering Committee, 1997-1998, 2013-2014
Editorial Boards
- The Journal of Immunology (section editor, 1993-1998)
- Current Protocols in Immunology (1989-date)
- Molecular Immunology (2002-date)
Special Interest Groups
Immunology, Structural Biology
Selected Publications
Ahmad J, Jiang J, Boyd LF, Zeher A, Huang R, Xia D, Natarajan K, Margulies DH. Structures of synthetic nanobody-SARS-CoV-2 receptor-binding domain complexes reveal distinct sites of interaction. J Biol Chem. 2021 Oct;297(4):101202.
Puig M, Ananthula S, Venna R, Kumar Polumuri S, Mattson E, Walker LM, Cardone M, Takahashi M, Su S, Boyd LF, Natarajan K, Abdoulaeva G, Wu WW, Roderiquez G, Hildebrand WH, Beaucage SL, Li Z, Margulies DH, Norcross MA. Alterations in the HLA-B*57:01 Immunopeptidome by Flucloxacillin and Immunogenicity of Drug-Haptenated Peptides. Front Immunol. 2021 Feb 9;11:629399.
Panda AK, Gangaplara A, Buszko M, Natarajan K, Boyd LF, Sharma S, Margulies DH, Shevach EM. Cutting Edge: Inhibition of the Interaction of NK Inhibitory Receptors with MHC Class I Augments Antiviral and Antitumor Immunity. J Immunol. 2020 Aug 1;205(3):567-572.
McShan AC, Natarajan K, Kumirov VK, Flores-Solis D, Jiang J, Badstübner M, Toor JS, Bagshaw CR, Kovrigin EL, Margulies DH, Sgourakis NG. Peptide exchange on MHC-I by TAPBPR is driven by a negative allostery release cycle. Nat Chem Biol. 2018 Aug;14(8):811-820. *
Jiang J, Natarajan K, Boyd LF, Morozov GI, Mage MG, Margulies DH. Crystal structure of a TAPBPR-MHC I complex reveals the mechanism of peptide editing in antigen presentation. Science. 2017 Nov 24;358(6366):1064-1068.
Natarajan K, McShan AC, Jiang J, Kumirov VK, Wang R, Zhao H, Schuck P, Tilahun ME, Boyd LF, Ying J, Bax A, Margulies DH, Sgourakis NG. An allosteric site in the T-cell receptor Cβ domain plays a critical signalling role. Nat Commun. 2017 May 16;8:15260.
Research Group
BMSBS studies the structural and molecular details of cellular recognition in immune response. We focus on cell surface molecules (MHC I and II, T cell receptors, NK cell receptors) using genetic, biochemical, biophysical tools including X-ray and cryo-EM to understand how crucial components interact. We study viral immunoevasins and how viral structures are recognized by antibodies.

