Lymphocyte Biology Section
Ronald N. Germain, M.D., Ph.D.
Chief, Laboratory of Immune System Biology
Chief, Lymphocyte Biology Section
Director, Center for Advanced Tissue Imaging (CAT-I)

Major Areas of Research
- Intravital imaging, analysis, and modeling of immune cell dynamics and in vivo activity
- Control of cell migration and cell-cell interactions by structural and chemical cues
- Multiplex imaging of cell phenotype, signaling, and function in complex tissues
- Systems-level analysis of immune cell signaling and responses to infection
- Human immune analysis using systems biology methods
Program Description
Past work and recent emphasis on applying imaging methods to analysis of the innate and adaptive immune systems
The Lymphocyte Biology Section (LBS) has made numerous contributions to the understanding of the cell biology of antigen processing and presentation by MHC class I and especially class II molecules. It also has examined recognition of these ligands by T cells with a focus on the signaling mechanisms involved in ligand discrimination. Since the early 2000’s, the LBS has conducted analysis of immune cell behavior in vivo using methods of intravital 2-photon imaging that it helped pioneer, providing real-time, high-resolution visualization of immune-cell dynamics in situ. More recently, the LBS has developed novel, highly multiplex section and volume imaging methods (Histo-cytometry, IBEX, and Ce3D) that allow an unprecedented analysis of cell phenotype, signaling, function, and location in complex tissue settings. These various imaging technologies are being used with more conventional molecular and cellular immunological methods to 1) describe the dynamics of innate and adaptive immune cell movement in lymphoid and non-lymphoid tissue; 2) localize the sites and duration of the cell-cell interactions involved in the development of adaptive immune responses; 3) analyze how differences in these aspects of cell migration and interaction affect differentiation events and functional immunity; and 4) investigate the dynamic behavior and effector activities of innate and adaptive immune cells in non-lymphoid sites.
Our studies have allowed us to determine how long a T cell spends in contact with an antigen-bearing dendritic cell, the role of both physical (the fibroblastic reticular cell network in lymph nodes) and chemical (chemokine) cues in controlling T-cell migration in secondary lymphoid tissues, the importance of SAP in differential attachment of T cells to antigen-presenting dendritic cells versus B cells and in germinal center formation, the signals guiding neutrophil migration in infected to damaged tissue settings, epithelial cell TLR control of dendritic-cell extension into the gut lumen for bacterial sampling, the interplay of myeloid and lymphoid cells within mycobacterial granulomas in the liver, the complex set of lymphocyte-dendritic cell interactions that underlie effective cell-mediated immune responses involving both CD4 and CD8 T cells, and the key role of resident tissue macrophages in preventing neutrophilic tissue damage. Videos from published papers reporting these findings are available in the Videos tab.
The static imaging methods developed in the LBS have contributed to a new understanding of how dendritic cell subsets are positioned within lymph nodes and the impact of this uneven distribution on functional biology post-infection or vaccination. They have also revealed the operation of regulatory T cells in small clusters within lymph nodes where conventional autoreactive cells produce IL-2 in response to self-antigens and are then silenced by TCR and IL-2 feedback activation of the associated Tregs. In the past few years, the Section has begun to apply these methods to an deeper understanding of the immune response to tumors, both in mouse models, and human material. Future work will emphasize both additional basic investigations and new studies of human material, such as samples from cancer patients receiving checkpoint inhibitor therapy. The expectation is that Histo-cytometry, IBEX, and Ce3D methods can provide critical new insight into why some and not other patients respond to these treatments, helping to assign individuals to optimal treatment groups and also aiding in development of new combination therapies for initial non-responders. Some of these studies will be conducted by a new Center for Advanced Tissue Imaging (CAT-I) jointly sponsored within the LBS by NIAID and the National Cancer Institute.
Imaging Immune Cell Dynamics and Function
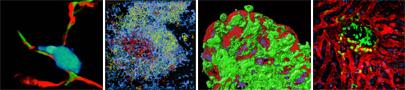
From L to R: Polarized T cell (cyan) associated with desmin+/ERTR-7+ (red and green) fibroblastic reticular cell (FRC) fibers; composite image of polyclonal B cells in a lymph node follicle (blue) and the migration tracks of wild-type (red) and SAP KO (green) T cells in the follicle and germinal center; dendritic cells (green) extending (‘balloon bodies”) into the gut lumen from beneath the villus epithelial cell layer (red); and activated T cells (green) in a BCG-induced liver granuloma surrounded by intact liver sinusoids (red blood tracer) and hepatocytes (blue nuclei).
AAI Excellence in Mentoring Award
Congratulations to Ronald N. Germain, M.D., Ph.D., Chief of the Laboratory of Immune System Biology, Chief of the Lymphocyte Biology Section, and Director of the Center for Advanced Tissue Imaging (CAT-I), for being awarded the 2025 American Association of Immunologists’ (AAI) Excellence in Mentoring Award. This prestigious award recognizes his outstanding career contributions to future generations of scientists and emphasizes the significance of the mentor-trainee relationship in the scientific community. Dr. Germain will receive the award during the AAI annual meeting in May. Read more about the AAI Excellence in Mentoring Award.
Biography
Education
M.D., Ph.D., 1976, Harvard Medical School, Harvard University
Sc.B., Sc.M., 1970, Brown University
Dr. Germain received his Sc.B. and Sc.M. from Brown University in 1970 and his M.D. and Ph.D. from Harvard Medical School and Harvard University in 1976. From 1976 to 1982, he served as an instructor, assistant professor, and associate professor of pathology at Harvard Medical School. From 1982 to 1987, he worked as a senior investigator in the Laboratory of Immunology (LI). In 1987, he was appointed Chief of the Lymphocyte Biology Section. In 1994, Dr. Germain was named Deputy Chief of LI. In 2006, he became director of the NIAID Program in Systems Immunology and Infectious Disease Modeling, which became the Laboratory of Systems Biology in 2011 and for which he served as Chief of the Laboratory. He also served as an associate director of the Trans-NIH Center for Human Immunology, Inflammation, and Autoimmunity (CHI) from 2010-2018. In 2018, the LI and LSB merged to create the Laboratory of Immune System Biology (LISB) with Dr. Germain as Chief of the Laboratory. Since receiving his doctoral degrees, he has led a laboratory investigating basic immunobiology. He and his colleagues have made key contributions to our understanding of MHC class II molecule structure–function relationships, the cell biology of antigen processing, and the molecular basis of T cell recognition.
More recently, his laboratory has explored the relationship between immune tissue organization and control of immunity studied using dynamic and static in situ microscopic methods that his laboratory helped pioneer. His group also conducts research in quantitative modeling of immune signaling circuits and in the broader application of the methods of systems biology to immunological questions. He has published more than 400 scholarly research papers and reviews and serves on the editorial boards of many scientific journals.
Among his numerous honors, he was elected as an associate (foreign) member of EMBO (2008), elected to the National Academy of Medicine (2013) and to the National Academy of Sciences (2016), received the Meritorious Career Award from the American Association of Immunologists (2015), was chosen as NIAID Outstanding Mentor (2016), named a Distinguished Fellow of the American Association of Immunologists (2019) and designated an NIH Distinguished Investigator. He has trained more than 70 postdoctoral fellows, many of whom hold senior academic and administrative positions at leading universities and medical schools.
Selected Publications
Liu Z, Gerner MY, Van Panhuys N, Levine AG, Rudensky AY, Germain RN. Immune homeostasis enforced by co-localized effector and regulatory T cells. Nature. 2015 Dec 10;528(7581):225-30.
Mao, K., Baptista, A.P., Tamoutounour, S., Zhuang, L., Bouladoux, N., Martins, A.J., Huang, Y., Gerner, M.Y., Belkaid, Y., and Germain, R.N. Innate and adaptive lymphocytes sequentially shape the gut microbiota and lipid metabolism. Nature 554:255-259, 2018.
Uderhardt, S., Martins, A.J., Tsang, J.S., Lämmermann, T., and Germain, R.N. Resident macrophages cloak tissue microlesions to prevent neutrophil-driven inflammatory damage. Cell 177:541-555, 2019.
Radtke, A.J., Kandov, E., Lowekamp, B., Speranza, E., Chu, C.J., Gola, A., Thakur, N., Shih, R., Yao, L., Yaniv, Z.R., Beuschel, R.T., Kabat, J. Croteau, J., Davis, J., Hernandez, J.M., and Germain, R.N. IBEX: A versatile multiplex optical imaging approach for deep phenotyping and spatial analysis of cells in complex tissues. Proc Natl Acad Sci U S A. 117:33455-33465, 2020.
Gola, A., Dorrington, M.G., Speranza, E., Sala, C., Shih, R.M., Radtke, A.J., Wong, H.S., Baptista, A.P., Hernandez, J.M., Castellani, G., Fraser, I.D.C., and Germain, R.N. Commensal-driven immune zonation of the liver promotes host defence. Nature. 589:131-136, 2021.
Wong, H.S., Park, K., Gola, A., Baptista, A..P, Miller, C.H., Deep, D., Lou, M., Boyd, L.F., Rudensky, A.Y., Savage, P.A., Altan-Bonnet, G., Tsang, J.S., and Germain, R.N. A local regulatory T cell feedback circuit maintains immune homeostasis by pruning self-activated T cells. Cell 184:3981-3997, 2021.
Videos from LBS Research
Movie 1: T cells exit HEVs in limited locations
A single z slice from an intravital four-dimensional data set showing numerous T cells (red) exiting HEV (green) in a lymph node via lucent areas that appear to be gaps in the FRC sheath (“exit ramps”). The playback speed is 300x for both the main and zoomed image. See Bajénoff M, Egen JG, Koo LY, Laugier JP, Brau F, Glaichenhaus N, Germain RN. Stromal cell networks regulate lymphocyte entry, migration, and territoriality in lymph nodes.Immunity. 2006 Dec;25(6):989-1001.
Movie 2: T cells migrate along the FRC network
Movie 3: Role of integrins in neutrophil migration and swarm formation
An intravital four-dimensional data set showing wild-type mouse neutrophils (red) swarming around a central point of localized tissue damage and the failure of talin-deficient (integrin-defective) neutrophils (green) to participate in swarm formation in a mouse ear. Both types of neutrophils migrate equivalently in the dermis, showing that high affinity integrin function is not necessary for such movement in dense tissue but is required for isolating the damage from surrounding viable tissue.
See Lämmermann T, Afonso PV, Angermann BR, Wang JM, Kastenmüller WK, Parent CA, Germain RN. Neutrophil swarms require LTB4 and integrins at sites of cell death in vivo. Nature. 2013 Jun 20;498(7454):371-375.
Movie 4: Neutrophils responding to a skin barrier breach
An intravital four-dimensional data set showing mouse neutrophils (green) leaving a blood vessel (blue) and migrating towards the site of a sand fly bit in the center of the imaging field. Red objects are genetically labeled Leishmania major organisms deposited by the sand fly.
See Peters, Nathan C, Jackson G Egen, Nagila Secundino, Alain Debrabant, Nicola Kimblin, Shaden Kamhawi, Phillip Lawyer, Michael P Fay, Ronald N Germain, and David Sacks. 2008. “In Vivo Imaging Reveals an Essential Role for Neutrophils in Leishmaniasis Transmitted by Sand Flies..” Science (New York, N.Y.) 321 (5891): 970–74. doi:10.1126/science.1159194.
Movie 5: Collateral tissue damage due to neutrophil swarming part 1
An intravital four-dimensional data set showing mouse neutrophils (red) and migrating towards the site of laser-induced tissue damage the center of the imaging field (white circle).
Movie 6: Collateral tissue damage due to neutrophil swarming part 2
Movie 7: Drainage and uptake of particulate antigens
Intravital 4D imaging dataset showing a mouse lymph node draining the site of vaccine particle inoculation the ear. Cyan =collagen in the capsule of the lymph node. Green = CD11c+ lymphatic sinus resident dendritic cells. Red = CD169+ lymphatic sinus resident macrophages. Blue = tracer in the draining lymph.
See Gerner, Michael Y, Parizad Torabi-Parizi, and Ronald N Germain. 2015. “Strategically Localized Dendritic Cells Promote Rapid T Cell Responses to Lymph-Borne Particulate Antigens..” Immunity 42 (1): 172–85. doi:10.1016/j.immuni.2014.12.024.
Movie 8: Direct capture of lymph draining particles by dendritic cell extending into the lymphatic sinuses
Intravital 4D imaging dataset zoon in showing CD11c+ dendritic ell capture of vaccine particles in a mouse lymph node draining the site of inoculation the ear. Green = CD11c+ lymphatic sinus resident dendritic cells. Yellow = vaccine particles. Blue = tracer in the draining lymph.
Movie 9: Three-dimensional reconstruction of DC extensions across the epithelial layer of the terminal ileum
Movie 10: Different shape of trans-epithelial DC extensions
Movie 11: Rapid association of blood-borne BCG with Kupffer cells observed in MHCII-EGFP mice
Movie 12: Visualizing tissues and immune cells in 3D
Animation of a 3D image of intact (non-sectioned) mouse lymph node imaged using the LBS-developed method of Ce3D clearing and staining. Cyan = B220 staining of B cells. Yellow = CD31 staining of vascular endothelial cells. White = LYVE-1 staining of lymphatic endothelial cells. Green = CD8 staining of T cells and CD8+ dendritic cells. Red = CD169 staining of macrophages. Image first shows vascular tree, then adds B cell follicles, then shows in silico computer sectioning of the entire lymph node.
Movie 13: Visualizing tissues and immune cells in 3D - Confetti animal
Animation of 3D image of mouse brain from a Confetti animal with multiple fluorescent proteins expressed in neurons using the LBS-developed method of Ce3D clearing.
Movie 14: Magnified view of a portion of Movie 13
Movie 15: Sequential migration of neutrophils from the epidermis/dermis into sites of sand fly proboscis penetration through the stratum corneum
Tools/Resources/Core Facilities
Center for Advanced Tissue Imaging (CAT-I): performs advanced imaging studies on human material in a collaborative manner for projects approved by a Scientific Advisory Board composed of NIAID and NCI scientists. CAT-I has the advanced microscopes , ancillary tissue processing equipment, and highly trained staff needed for these studies.
Affiliations
Research Networks
Chan-Zuckerberg Initiative:
- Seed Networks for Tissue Analysis as part of the Human Cell Atlas framework
- Inflammation Network
Training Programs
NIG – UPenn PhD Program
Research Group
LBS has made numerous contributions to understanding cell biology of antigen processing and presentation by MHC class I and especially class II molecules. It also has examined recognition of these ligands by T cells with a focus on the signaling mechanisms involved in ligand discrimination. Various imaging technologies are used with more conventional molecular and cellular immunological methods.


