A healthy volunteer receives an intravenous infusion of mAb114 in a Phase 1 clinical trial held at the NIH Clinical Center in Bethesda, Maryland (May 21, 2018).
Discovery of Therapeutic Antibody for Ebola
Building on years of association with physician-scientists from the National Institute of Biomedical Research (INRB) in the Democratic Republic of the Congo (DRC), VRC scientists identified a survivor of the 1995 Kikwit Ebola outbreak, who long after infection maintained a strong antibody response against the Ebola virus. In collaboration with the INRB and Institute for Biomedical Research in Switzerland, the team defined the structural basis of neutralization for these antibodies leading to the discovery of mAb114, a monoclonal antibody that binds to the receptor-binding domain. Preclinical studies of mAb114 demonstrated promising results and signaled a potential therapeutic application for the biologic. In early phase clinical trials, mAb114 was found to be safe and well-tolerated following intravenous infusion. In a randomized controlled trial of investigational Ebola treatments conducted in an outbreak setting, mAb114 was shown to significantly reduce mortality from Ebola virus disease. Developed by Ridgeback Biotherapeutics, mAb114 was approved by the Food and Drug Administration (FDA) in 2020 as EbangaTM (ansuvimab-zykl).
Related Publications
FDA approves treatment for ebola virus, FDA Press Release. Dec 21, 2020.
A Randomized, Controlled Trial of Ebola Virus Disease Therapeutics, N Engl J Med. 2019 Dec 12;381(24):2293-2303.
Safety, tolerability, pharmacokinetics, and immunogenicity of the therapeutic monoclonal antibody mAb114 targeting Ebola virus glycoprotein (VRC 608): an open-label phase 1 study, Lancet. 2019 Mar 2;393(10174):889-898.
Protective monotherapy against lethal Ebola virus infection by a potently neutralizing antibody, Science. 2016 Mar 18;351(6279):1339-42.
Structural and molecular basis for Ebola virus neutralization by protective human antibodies, Science. 2016 Mar 18;351(6279):1343-6.
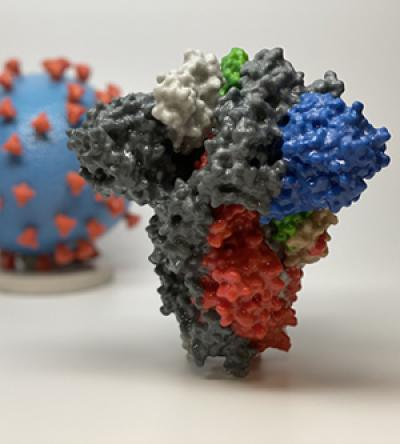
3D print of a SARS-CoV-2 spike protein in front of a SARS-CoV-2 virus particle 3D print.
Rapid Design of COVID-19 Vaccine Candidates
Prior to the identification of SARS-CoV-2, VRC research scientists had demonstrated that a stabilized spike protein antigen for betacoronaviruses could be a key target for vaccines, therapeutics, and diagnostics with research on SARS, HKU1, and MERS. When COVID-19 emerged, the VRC team, with collaborators from the University of Texas at Austin, were promptly able to identify the relevant two proline mutation (2P) for spike protein stabilization. The favorable safety and immunogenicity profile demonstrated in early trials of vaccine candidates encoding the S2P design across vaccine platforms, allowed several COVID-19 vaccine products to advance into large-scale efficacy trials. In less than one year from pathogen emergence, multiple vaccines incorporating the S2P design of the spike antigen were approved for emergency use by both national and international regulators.
Related Publications
SARS-CoV-2 mRNA vaccine design enabled by prototype pathogen preparedness, Nature. 2020 Oct;586(7830):567-571.
Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation, Science. 2020 Mar 13;367(6483):1260-1263.
Immunogenicity and structures of a rationally designed prefusion MERS-CoV spike antigen, Proc Natl Acad Sci USA. 2017 Aug 29;114(35):E7348-E7357.
Pre-fusion structure of a human coronavirus spike protein, Nature. 2016 Mar 3;531(7592):118-21.
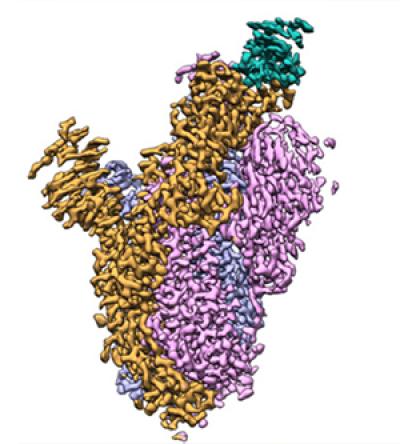
A high-resolution cryo-EM map of the LY-CoV555-spike complex.
Discovery of COVID-19 Therapeutic Antibody
Immediately following the emergence of SARS-CoV-2, VRC scientists applied their extensive experience in monoclonal antibody (mAb) discovery to identify a therapeutic mAb for COVID-19. Working in collaboration with AbCellera, VRC researchers isolated and characterized a neutralizing antibody derived from a COVID-19 convalescent blood sample. When evaluated as a therapeutic strategy in a clinical setting by development partner Lilly, the monoclonal antibody termed LY-CoV555, demonstrated efficacy. LY-CoV555 was the first mAb to be issued emergency use authorization by the Food and Drug Administration (FDA) for COVID-19 treatment as bamlanivimab.
Related Publications
Fact sheet for health care providers emergency use authorization (EUA) of bamlanivimab and etesevimab, FDA Fact Sheet
The neutralizing antibody, LY-CoV555, protects against SARS-CoV-2 infection in nonhuman primates, Sci Transl Med. 2021 May 12;13(593):eabf1906.
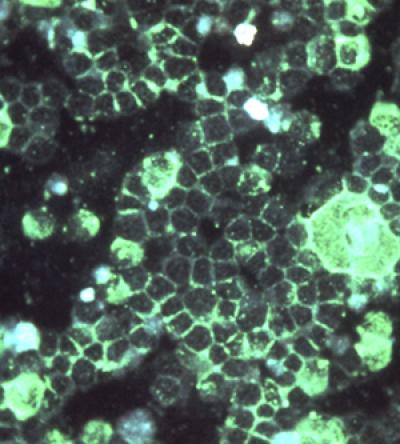
Using indirect immunofluorescence microscopy, this photomicrograph revealed the presence of the respiratory syncytial virus (RSV) in an unidentified tissue sample. RSV is the most common cause of bronchiolitis and pneumonia among infants and children under 1 year of age.
Structure-Based Design to Advance RSV Vaccine Candidates
After years of research into the structural and antigenic properties of respiratory syncytial virus (RSV), VRC scientists made a critical breakthrough by defining the atomic-level structure of an RSV prefusion protein. This discovery led to the development of vaccine candidate DS-Cav1, which introduced stabilizing disulfide (DS) and cavity filling (Cav) mutations to generate an immunogen. With promising preclinical results, the vaccine candidate was advanced into phase 1 study and demonstrated a significant boost in neutralizing activity. This study established a clinical proof of concept for structure-based vaccine design, a novel approach to immunization against viral diseases. Products based on this work are in advanced clinical trials with promising results shown from ongoing Phase 2 and Phase 3 studies, a first in the RSV field.
Related Publications
Safety, tolerability, and immunogenicity of the respiratory syncytial virus prefusion F subunit vaccine DS-Cav1: a phase 1, randomised, open-label, dose-escalation clinical trial, Lancet Respir Med. 2021 Oct;9(10):1111-1120.
Vaccination with prefusion-stabilized respiratory syncytial virus fusion protein induces genetically and antigenically diverse antibody responses, Immunity. 2021 Apr 13;54(4):769-780.e6.
A proof of concept for structure-based vaccine design targeting RSV in humans, Science. 2019 Aug 2;365(6452):505-509.
Structure-based design of a fusion glycoprotein vaccine for respiratory syncytial virus, Science. 2013 Nov 1;342(6158):592-8.
Structure of RSV fusion glycoprotein trimer bound to a prefusion-specific neutralizing antibody, Science. 2013 May 31;340(6136):1113-7.

